West Nile Virus is an arbovirus of the flavivirus genus which is able to infect a range of vertebrate hosts such as humans, birds and horses1. Its history in the eastern hemisphere dates back to 1937 and is carried by Culex species of mosquitoes, which can infect mammals via bites1. This arbovirus was introduced to the Western hemisphere in 1999 in New York City (NYC) where it infected both birds and humans. While public health officials confirmed 61 human cases across NYC, door to door serostudies conducted throughout the next year estimated the infection rate to be up to 2.6% in the Queens area. This infection rate would estimate that there were closer to 8200 cases in Queens, most of which went undiagnosed due to asymptomatic infections1.
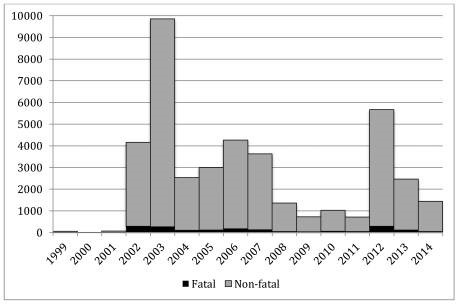
Many public health officials believed the introduction of WNV in the U.S. would spread slowly outside of NYC as Culex mosquitoes, the carrier of WNV, increased their range. However, in 2002, it was shown that WNV infected mosquitoes had rapidly disseminated when an outbreak of WNV was identified in 28 states with 4,156 confirmed cases of WNV2,3. Then, in 2003, the Culex mosquitoes carrying WNV were confirmed in all Eastern and Midwest states (sans Maine) and infected 9,862 individuals in the U.S. alone1,2 (Figure 1).

While mild infection of WNV and other arboviruses are similar to a seasonal cold in clinical symptoms, approximately 1 in 150 individuals will develop neuroinvasive disease which can lead to convulsions, organ failure and death3,4. Neuroinvasive disease became a reportable illness in the U.S. in 2001, while non-neuroinvasive disease became reportable in 2004, with all states having their own surveillance systems which then report to the national network called arboNET, which is maintained by the Centers for Disease Control and Prevention (CDC)4. Furthermore, recent research suggests that many arboviruses can lead to neurological deficits and diseases such as Guillain-Barre syndrome or congenital diseases such as microcephaly during pregnancy later on in life after initial infection4.
Mitigation of WNV is of concern to health officials due to the risk of developing the severe symptoms which can lead to death and the compounded concern of neurological deficits resulting from infection. Arboviruses also tend to have a cyclical infection cycle where a specific virus will have an increased number of infections during one year, level out over the next 8-10 years and then peak again2,4.
There is no treatment for West Nile and hospitals and providers generally provide care to cover the symptoms. Similarly, no vaccines in clinical trials been successful enough to be mass-produced and distributed4,5. With no other treatment options and the cyclical cycle of infection-causing different arboviruses to peak in different years, a non-disease specific way of controlling the vector (mosquito) populations is currently the supported means of controlling arbovirus infection rates (Figure 2). Both passive and active forms of vector control will be evaluated for efficiency over the last decade, with a recommendation towards using genetically modified mosquitoes within integrated vector management (IVM) approach as a long term means of vector control.
Epidemiology
Since its introduction to NYC in 1999, WNV has become the leading arbovirus infection and encephalitis in the U.S. Only about 1 in 5 individuals with an infection will have symptoms which include what is defined as an acute system febrile illness, the other 4 out of 5 individuals will be asymptomatic4. The most common symptoms seen with systemic febrile illness are a headache, arthralgia, rash, and gastrointestinal disturbances. In 1 out of 150 cases, individuals will develop neuroinvasive diseases such as meningoencephalitis, meningitis, encephalitis or acute flaccid paralysis (AFP), any one of which can be life threatening3,4.
Like many illnesses, WNV infection can affect vulnerable populations differently and a paper by CDC in 2010 calculated the rates of infection for different populations after analyzing the 28,961 cases that met clinical and laboratory criteria for WNV infection between 1999-2008. Of these cases, 17,139 (approximately 59%) were defined as non-neuroinvasive and 11,822 (41%) were defined as neuroinvasive (Table 1).

Out of the 17,139 cases reported of non-neuroinvasive, males accounted for 8,972 approximately, 52% of the confirmed cases (Table 1). Similarly, for neuroinvasive cases, males accounted for 6,887 cases out of 11,822 reported cases (Table 1). Neuroinvasive disease and non-neuroinvasive affected older populations at a higher rate, with the median age being 57 years old and 47 years old, respectively. In Neuroinvasive, nearly 46% of cases were 60 years old or above, while 45% of non-neuroinvasive were between 40-59 years old. For both forms of the disease, between 88-95% of those affected were Caucasian with 82-90% of cases being non-Hispanic (Table 1).
Out of the 8925 cases with hospital data, the non-neuroinvasive cases had a hospitalization rate of 20.6/1000 and a fatality rate of 0.96/1000. Of the fatal cases, being above 70 years old increased your risk as 85 of 86 fatal cases were above 70. Neuroinvasive has a higher rate of hospitalization going from 20.6/1000 in non-neuroinvasive to 86/1000 in neuroinvasive1,4. Neuroinvasive cases also results in a higher fatality rate going from 47/1000 in non-neuroinvasive to 178/1000 in neuroinvasive. The mean annual incidence of neuroinvasive cases rose from 5/1000 for those younger than 10 to 135/1000 for those above 70. This trend continues for gender differences and age with a mean incidence of 48/1000 in men and 33/1000 for women below 70 years old but increasing to 200/1000 in men and 89/1000 for women above 70 years old1,4.
West Nile Virus is an arbovirus which is transmitted by mosquitoes through mosquito bites. For both forms of the disease, between 91-94% of cases occurred between July and September, the peak mosquito season4. As you can see in Figure 1, WNV disseminated across the U.S. within 6 years, and by 2006 is found in every state in the continental U.S. except for Maine. Similarly, all arboviruses follow a cyclical pattern of infection where one virus will surge during one year while the rest maintain a low infection rate and then the surging virus will level out for 8-10 years while other viruses surge periodically. Then after 8-10 the first virus will surge again; for West Nile virus you see this trend in 2002-2003 when it first enters the United States, levels out between 2003-2011 and then surges again in 2012 (Figure 2).
References
- Assessing Capacity for Surveillance, Prevention, and Control of West Nile Virus Infection—United States, 1999 and 2004. Jama. 2006;295(15):1765.
- Provisional Surveillance Summary of the West Nile Virus Epidemic—United States, January–November 2002. Jama. 2003;289(3):293.
- Roehrig J. West Nile Virus in the United States — A Historical Perspective. Viruses. 2013 Oct;5(12):3088–108.
- Patel H, Sander B, Nelder MP. Long-term sequelae of West Nile virus-related illness: a systematic review. The Lancet Infectious Diseases. 2015;15(8):951–9.
- Bellini R, Zeller H, Bortel WV. A review of the vector management methods to prevent and control outbreaks of West Nile virus infection and the challenge for Europe. Parasites & Vectors. 2014;7(1):323.
- Benelli G. Research in mosquito control: current challenges for a brighter future. Parasitology Research. 2015;114(8):2801–5.
- Wilke ABB, Marrelli MT. Paratransgenesis: a promising new strategy for mosquito vector control. Parasites & Vectors. 2015;8(1).
- Dodson BL, Hughes GL, Paul O, Matacchiero AC, Kramer LD, Rasgon JL. Wolbachia Enhances West Nile Virus (WNV) Infection in the Mosquito Culex tarsalis. PLoS Neglected Tropical Diseases. 2014 Oct;8(7).
- Fradin MS, Day JF. Comparative Efficacy of Insect Repellents against Mosquito Bites. New England Journal of Medicine. 2002 Apr;347(1):13–8.
- Knox TB, Juma EO, Ochomo EO, Jamet HP, Ndungo L, Chege P, et al. An online tool for mapping insecticide resistance in major Anopheles vectors of human malaria parasites and review of resistance status for the Afrotropical region. Parasites & Vectors. 2014;7(1):76.
- Hamer GL, Anderson TK, Donovan DJ, Brawn JD, Krebs BL, Gardner AM, et al. Dispersal of Adult Culex Mosquitoes in an Urban West Nile Virus Hotspot: A Mark-Capture Study Incorporating Stable Isotope Enrichment of Natural Larval Habitats. PLoS Neglected Tropical Diseases. 2014;8(3).
- Jost CA, Pierson TC. Antibody-Mediated Neutralization of West Nile Virus: Factors that Govern Neutralization Potency. West Nile Encephalitis Virus Infection. 2009;219–47.






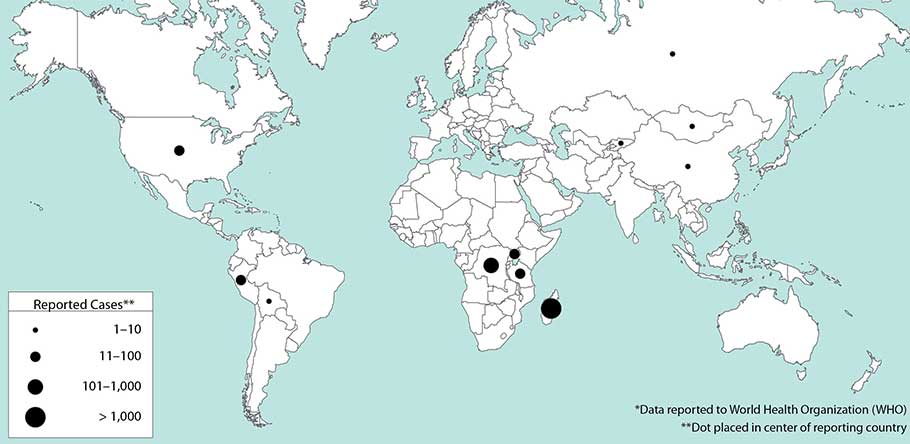


 Bogoch et. Al. 2016. Anticipating the international spread of Zika virus from Brazil. The Lancet.
Bogoch et. Al. 2016. Anticipating the international spread of Zika virus from Brazil. The Lancet.